Watch our interview with Ineke Knot on our YouTube channel.
Ineke Knot is bringing local health monitoring of great apes into the genomic age. Over the course of the last four years, she has developed methods to identify different types of nematodes using a fully portable system. Her research has recently been published in an open access paper: DNA Barcoding of Nematodes Using the MinION.
Protecting Great Apes from Infections
As our close genetic relatives, primates are susceptible to human infections. When primate communities are in contact with humans – either local people, researchers or ecotourists – they are vulnerable to the transmission of infections. Infectious diseases can decimate local populations of great apes. In 2006, an outbreak of Ebola led to a gorilla population decline of about 83%.
“Humans and great apes are so closely genetically related, that a lot of our diseases that we carry can also be transmitted to great apes.”
Ineke Knot
Like humans, apes are social animals. But as Ineke points out, how can you ask a gorilla to practice social distancing? This makes real time health monitoring incredibly important, so that other measures can be implemented to protect the primates.
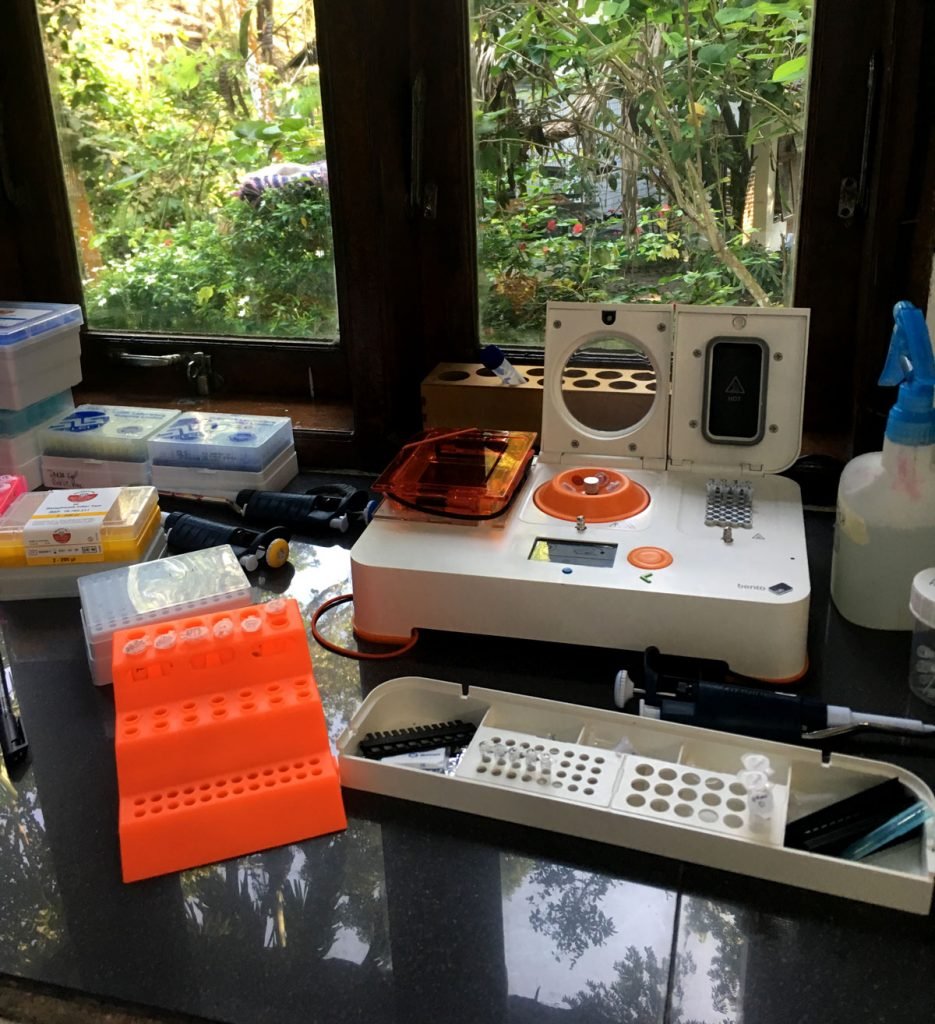

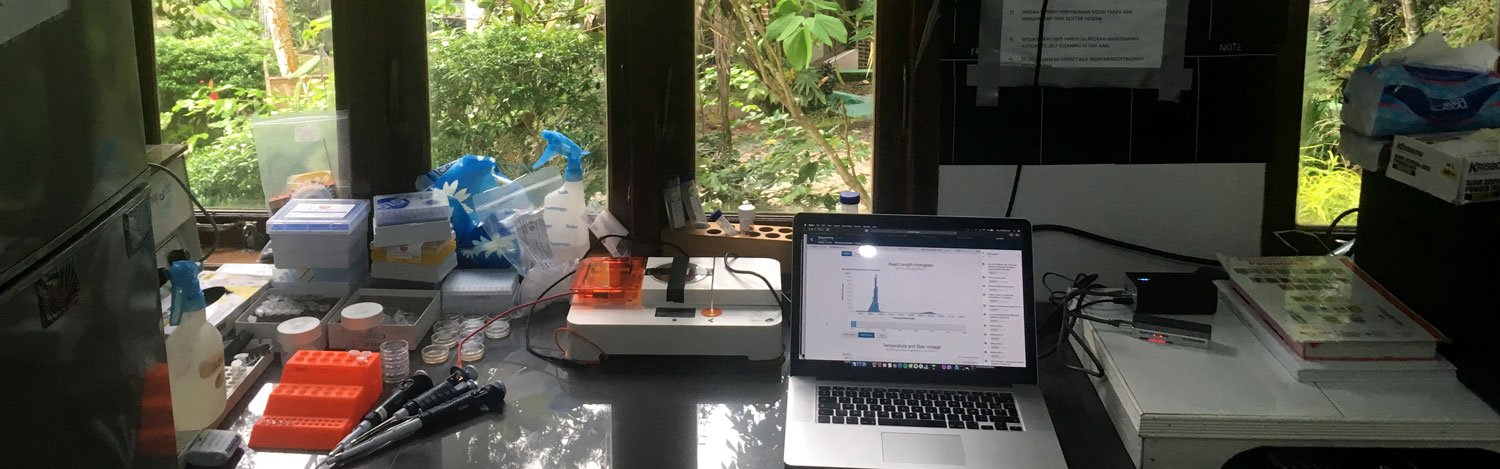
Temporary lab set-up at the veterinary clinic. Photo by Ineke Knot.
Using Portable Genomic Tools to Identify Nematodes
As part of a PhD with the University of Amsterdam, Ineke has been developing DNA-based field methods to identify parasites in great apes. Her research focuses on nematodes – microscopic worms that include several parasitic species.
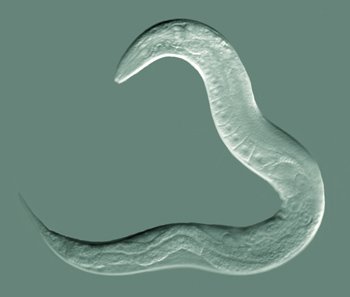
C. Elegans, one of the many types of nematodes. Photo: Bob Goldstein, UNC Chapel Hill
Traditionally, most zoonotic studies focus on viruses. Viral RNA evolves rapidly, and allows researchers to compare changes between a single generation. But working with viruses is notoriously difficult. RNA is more unstable and susceptible to degradation than DNA, which poses extra challenges when working in the field.
Interested in DNA methods and workflows? Subscribe for monthly insights.
To demonstrate proof of concept, Ineke has focused her attention on nematodes to apply portable genomics to conservation.
When I started my PhD, I realized that nematodes are a kind of neglected tropical disease. In a lot of areas of the world, parasite infections are still really widespread health problems.
Ineke Knot
Ineke ran tests with four different species, Anisakis simplex, Panagrellus redivivus, Turbatrix aceti, and Caenorhabditis elegans.
To identify the species with DNA barcoding, the team chose a common barcode for nematode species identification: a gene fragment from the 18S ribosomal RNA gene that is about ∼900 base pairs in length. This test could be applied to samples from a range of different environments, from faeces, to soil, to marine sediments.
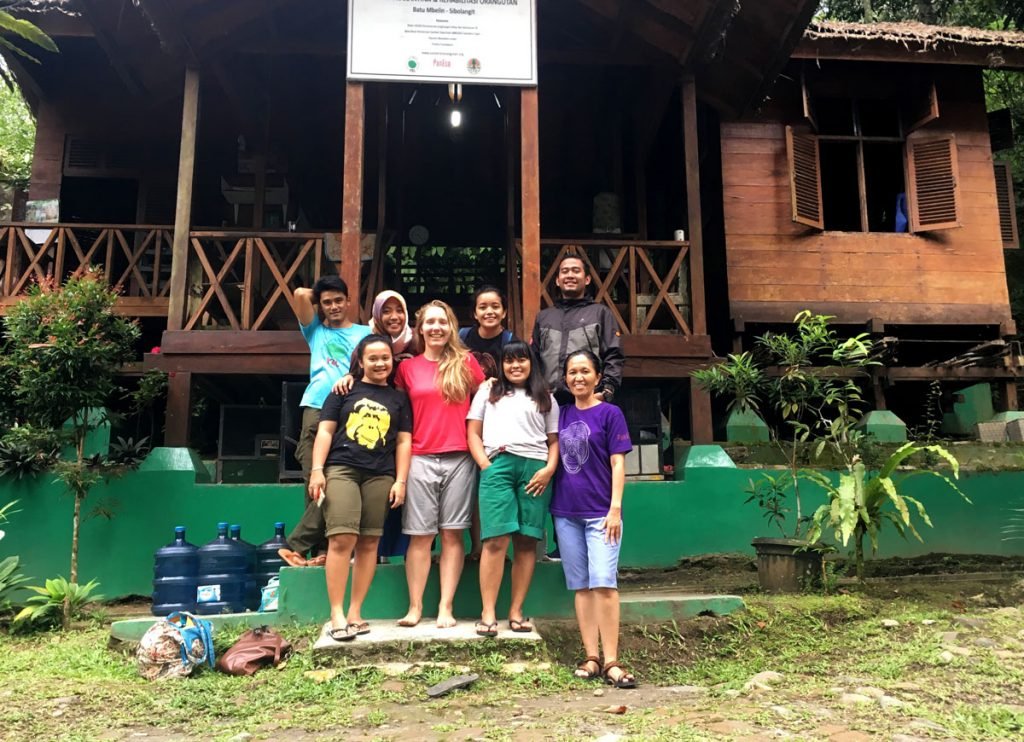

Ineke and the Sumatran Orangutan Conservation Programme team. Photo by Ineke Knot.
Sumatran Orangutan Conservation Programme (SOCP)
Last year, Ineke travelled to Indonesia to share her knowledge with the Sumatran Orangutan Conservation Programme. The NGO focuses on reintroducing orangutans to Bukit Tigapuluh National Park, an Indonesian forest where the species had been missing for over 150 years. As part of their rescue and rehabilitation work, the programme hopes to start using portable genomic tools like Bento Lab to study the microbiome of orangutans in their care.
Right now, the SOCP is fundraising in order to protect the orangutans from Covid19. If you are interested in helping, you can find more information on their fundraising website.
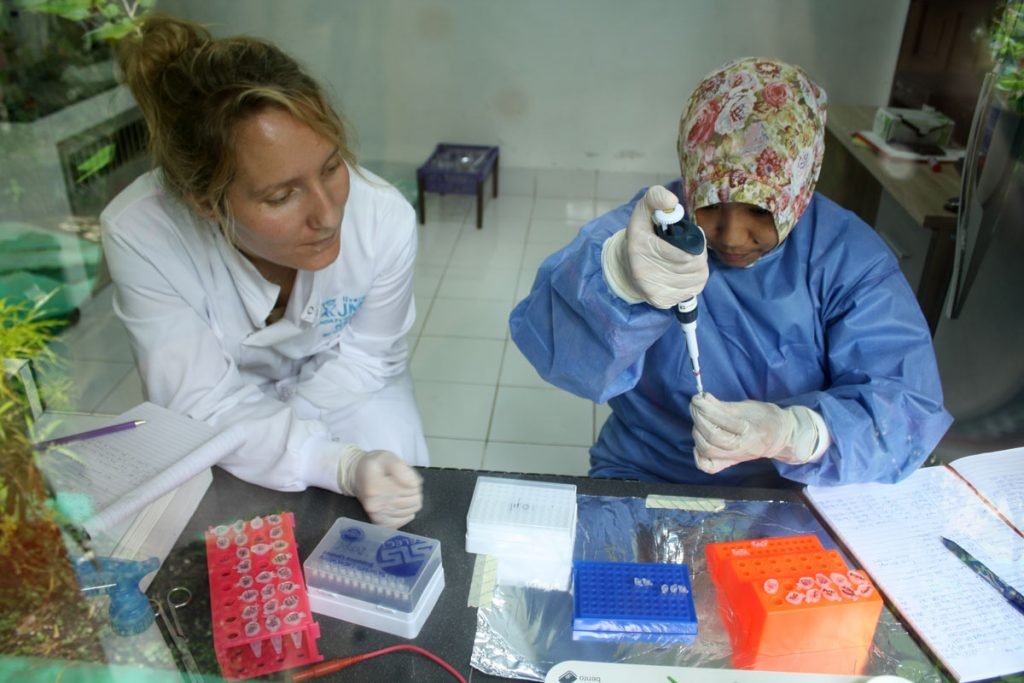


Training Local Staff and Spreading Knowledge
Providing training for the team in Sumatra has changed Ineke’s perspective. She is now on a mission to share her knowledge, and support wildlife conservationists that want to undertake molecular biology on remote field sites, like the Sumatran Orangutan Conservation Programme.
Looking for advice on using Bento Lab?
Book a free consultation or ask a question.If you are interested to learn more about using portable genomics in the field, Ineke will be running a free tutorial on 13th August with WILDLABS, a community platform for conservationists and technology experts.
You can keep up with Ineke by following her personal blog, and her Instagram. Ineke also talks about science on Twitter as @ieknot.
Hooray! It’s here: my first first-author publication! DNA barcoding of nematodes using the @nanopore #MinION and @theBentoLab. It’s #OpenAccess, follow the link to read the full article. If you’re interested in a summary, keep reading this thread. https://t.co/ww9tgXbgzF 1/12
— Ineke Knot (@ieknot) May 9, 2020