Brought to you by the team at Bento Bio on the mission to make DNA lab work accessible to all. We create powerful, easy-to-use devices, affordable supplies, and guides, to help anyone—from beginners to professionals—harness the power of PCR anywhere.
In this article we discuss some of the basics of using controls for experiments involving DNA extraction and PCR.
This article is intended to provide an overview, examples, and quick lookup tables, for complete beginners and experienced users alike.
You may find this to be useful information when planning experiments before you start, or when troubleshooting PCR workflows when things don’t work as expected.
By the end of this article you will have learned, or refreshed:
- What controls are
- Why controls are necessary in scientific experiments
- Why controls are used in DNA extraction and PCR
- The most common kinds of controls in PCR workflows
- How to make PCR controls and DNA extraction controls,
- How to interpret results when using controls
Looking for a compact PCR setup for your lab or extra equipment capacity at an affordable cost?
Bento Lab is a portable laboratory that combines 3 key tools — a thermal cycler for PCR, a gel electrophoresis unit, and a mini centrifuge — into a compact system for less than $2000.
Background to Controls
When running scientific experiments, it is important to design your experiments in ways that allow you to measure effects, test whether processes are working, and to troubleshoot when things go wrong. The best way to do this is to use control variables and control experiments.
Control variables are measurable elements of the experiment that are kept the same between experimental iterations. By changing only one variable and keeping the rest controlled, confounding or complicating factors are minimised and a clear link can be inferred between the tested variable and the result.
Control experiments are iterations of the larger experiment (wholly or in part) where a known experimental component is used to test part of the experimental process, or used to establish a baseline against which to compare the results of other experiments.
To do control experiments it is usually necessary to have experimental components of a type comparable to the subject of an experiment, and these should be of a known or default (i.e. “controlled”) state. These components can include sites, animals, plants, tissue samples, or DNA extracts. The control components can be individual or grouped, for example a single control plant or a control group of plants.
When controls are samples, a good control sample will be abundant enough to be used repeatedly as a control without exhausting the material, and be known to work well in the experimental process to produce a reliable result each time.
Interested in DNA methods and workflows? Subscribe for monthly insights.
General Experimental Controls in PCR-based applications
Before considering PCR-specific controls, it is useful to note that you should also plan in controls for your larger experiments overall – they do not just apply to PCR-based work!
If your experiment is not well designed and does not control for confounding factors then your DNA extractions and PCRs may be successful, but the overall experiment may be flawed or contain confounding effects.
When planning a complicated experiment it is therefore important to get expert advice on good experimental design and analysis of results, including on what controls to design into the experiment.
PCR Specific Controls in PCR-based applications
Controls are important to include in your DNA extraction and PCR processes to ensure that all the processes are working as required. This is because the processes from DNA extraction to gel electrophoresis will involve many steps, and problems can occur at each step. If problems occur then you will need to troubleshoot each step to find the problem.
Controls are also useful when optimising DNA extraction and PCR protocols, in which each step is systematically adjusted one at a time to improve results while keeping all the other processes the same. Since the results of a PCR application are only seen at the very end of the process (for example on an electrophoresis gel), when troubleshooting using controls it can be useful to work backwards through your workflow from PCR to DNA template dilution to DNA extraction to sample quality.
The main kinds of controls used in PCR applications are:
- Negative and positive PCR controls.
- Negative and positive DNA extraction controls
- Internal PCR assay controls
Negative PCR controls
Negative PCR controls allow you to test whether you have some form of contamination in your PCR workflow. They consist of PCRs where PCR grade water is added instead of DNA template.
Knowing that your PCRs are free from contamination is essential because otherwise all of your results could be caused by contamination and could be invalid. In some cases you may be able to differentiate between genuine results and weak contamination by amplicon size or by band brightness, but in many other cases it will not be so clear. In any case, at the first sign of PCR contamination you should carefully decontaminate your workflow and check your reagents for contamination.
Looking for advice on using Bento Lab?
Book a free consultation or ask a question.When using negative PCR controls:
- A negative result of no amplicon usually indicates that there is no systemic contamination in your workflow, unless all your PCRs failed.
- A positive result of an amplicon indicates that there is some systemic contamination in your PCR, from contaminated reagents, equipment, or your workspace.
It is important to note that if you see no positive PCR results at all then you have no idea if your negative PCR control worked properly or not. It could be possible that all PCRs failed!
In such cases using a negative PCR control and a positive PCR control would be needed to find out if the PCR is working or if it is failing.
The inferences that you can make from negative controls are shown in the table below:
| Sample PCR result | Negative PCR Control | Inferences and next steps |
|---|---|---|
| Amplicons observed | Not included | The PCR worked but might be contaminated. Repeat the experiment with a negative control. |
| Amplicons observed | Negative | The PCR worked and is unlikely to be contaminated. |
| No amplicons observed | Negative | The PCR may have worked (if testing for presence or absence), or may have failed (if a result was expected). Repeat the experiment with a positive PCR control to confirm for a genuine absence of the target, and if an amplicon was expected then also use a positive DNA extraction control to distinguish between PCR failure and DNA extraction failure. |
| Amplicons either observed or not observed | Positive | The PCR worked but has some form of systemic contamination It may be difficult to distinguish successful PCR products from PCR contaminants. |
Positive PCR controls
Positive PCR control experiments allow you to test whether your PCR is working in general, even if your sample PCRs fail. They consist of a sample of DNA template from an appropriate sample that has previously been shown to work well. They are best used in combination with a negative PCR control.
A positive result of an amplicon produced of the correct size indicates that the PCR in general worked, even if no other samples produced an amplicon.
A negative result of no amplicon being produced indicates that the PCR in general failed and needs troubleshooting.
Positive PCR controls are not always essential in routine PCR because any successful PCR within your PCR run will also demonstrate that the PCR in general is working. However, if all PCRs fail then you will need a positive PCR control to know whether the problem is your PCR or an issue with DNA extraction.
The inferences that you can make from using a negative control and a positive control together are shown in the table below:
| PCR result | Negative PCR control | Positive PCR control | Inferences and next steps |
|---|---|---|---|
| Amplicon observed | Negative | Positive | The PCR worked in general and for your samples, and is unlikely to be contaminated |
| Amplicon observed | Negative | Negative | The PCR worked in general and for your samples, and is unlikely to be contaminated. Your positive control needs troubleshooting or replacing. |
| Amplicons observed | Positive | Positive | The PCR in general worked and there is systemic contamination present. It may be difficult or impossible to distinguish successful sample PCRs from contamination. |
| Amplicons observed | Positive | Negative | The PCR in general worked. Double check that you put the controls in the correct wells since they could be reversed. If they were not reversed then there is systemic contamination present; it may be difficult or impossible to distinguish successful sample PCRs from contamination; and your positive control is not working and needs replacing. |
| No amplicons observed | Negative | Positive | The PCR in general worked and your samples failed. Troubleshoot your DNA extractions. |
| No amplicons observed | Negative | Negative | The PCR in general failed |
| No amplicons observed | Positive | Positive | The PCR in general worked and there is systemic PCR contamination. The sample PCRs all failed. Troubleshoot your DNA extractions. |
| No amplicons observed | Positive | Negative | The PCR in general worked and there is systemic PCR contamination as the only amplicon. Check that you didn’t reverse the control order as these are both opposite to the expected result. If you did not then the sample PCRs all failed. Troubleshoot DNA extraction. Your positive control is not working and needs replacing. |
How to make a positive PCR control
It is very useful to obtain or produce a positive PCR control when you start doing your experiments in case your PCRs fail.
Positive PCR controls usually consist of a known working DNA extract, such as a purified or crude DNA extract. A purified DNA extract should have a much longer storage life than a crude DNA extract.
When selecting a sample to use for a positive PCR control it can be useful to use an abundant DNA source from which it is very easy to extract DNA. If the PCR will be sequenced then it is also useful to select something that you would never expect to get in your sequencing results — this will aid detection of any contamination from the control in the sequencing results.
Example: If you are barcoding fungi then a button mushroom from a shop may be a good positive control. It is easy to source, can be dried in a good state, and has plenty of material. But if you are working on this genus (Agaricus) then a different type of fungus may be better as a control because any cross contamination will be much easier to detect.
Example: If you are sexing birds using DNA then DNA extracted from a small piece of chicken meat may provide a good positive control. It is easy to source and DNA extraction will be much easier than if using feathers or bloodspots.
A highly diluted PCR product can also be used as a convenient positive PCR control, for example a 10,000x dilution of a successful PCR product in TE buffer (for long term storage) or PCR grade water (for immediate use). Unfortunately, diluted PCR products do have some problems:
- They can be less stable than purified genomic DNA extracts
- They have an increased risk of producing primer dimers or non-specific amplification because past amplification may have already amplified trace amounts
- They can constitute a possible PCR contamination risk if not handled carefully in the pre-PCR stage
Looking for advice on using Bento Lab?
Book a free consultation or ask a question.Positive and negative DNA extraction controls
A positive DNA extraction control allows you to test whether there are issues with your DNA extraction method or whether your samples are of poor quality. It can be run while testing or optimising DNA extraction methods, or if your PCRs are not working and you have ruled out PCR failure as a cause. It is best used in combination with a negative DNA extraction control to test for DNA extraction contamination, and positive and negative PCR controls.
A negative DNA extraction control can be used to test whether there is contamination in any of your DNA extraction reagents or workflows. A negative DNA extraction control consists of working through the DNA extraction process without any material being added, and then using the same amount of end-product (which should lack any DNA) in place of DNA template in your PCR. The expected result is that a PCR produced with it is blank. If it is positive, and your PCR negative control is blank, then you have some kind of contamination in your DNA extraction workflow.
The inferences that you can make using positive and negative DNA extraction controls are shown in the table below:
| PCR result of new DNA extractions | Positive PCR control | Negative PCR control | Positive DNA extraction control | Negative DNA extraction control | Inferences and next steps |
|---|---|---|---|---|---|
| Strong amplicons observed | Positive | Negative | Positive | Negative | Your DNA extraction method was able to produce amplifiable DNA from your sample. Your sample is probably good quality. Your DNA extraction method is not contaminated. |
| Faint amplicons observed | Positive | Negative | Positive | Negative | Your DNA extraction method was able to produce amplifiable DNA from your sample but only a small amount. Your samples may be degraded. Your DNA extraction method is not contaminated. |
| No amplicons observed | Positive | Negative | Positive | Negative | Your DNA extraction method could not produce amplifiable DNA from your sample. Your samples may be degraded or you may be extracting too many PCR inhibitors. Try to dilute your PCR products 1:10 or 1:100. If this fails then your samples are either too poor quality or you need a different DNA extraction method for this sample type. |
| Any result | Any negative result | Any negative result | Any negative result | Any negative result | If you have any issues with your DNA extraction and PCR controls you should troubleshoot these and get them working before you can draw conclusions. |
How to make a positive DNA extraction control
A positive DNA extraction control should be a sample that is known to produce good quality DNA when extracted with your normal methods for this sample type.
It should be as similar to your experimental samples as possible but of high quality, for example a fresh collection, or a sample collected and frozen in good condition.
It should also be of sufficient quantity or easy to source so that you can use it for many experiments and in different extraction quantities.
Example: When DNA barcoding lichens, it is useful to have a standard large dried sample from which you have previously successfully extracted and amplified DNA to use as a control.
Example: When sexing birds, it is useful to have a set of reasonably fresh feathers from a single bird, or a set of blood spots from a single bird, from which you have previously extracted and amplified DNA to use as a control.
Internal PCR controls
For some PCR assays you may want to include an internal control so that you can show that each PCR contains amplifiable DNA, even if the main target DNA region of the assay is not present. Otherwise there would be no way of distinguishing a failed PCR for a sample from a lack of that target being present.
For PCR assays producing multiple bands, there is sometimes a risk that not all bands amplify because of degraded DNA or because of low concentrations of DNA. Internal controls can be used to highlight such reactions by acting as an amplicon that is more difficult to amplify (due to greater length) than the largest target amplicon. If the internal control fails then the results are inconclusive.
Internal controls are particularly important in assays where you need to know samples worked successfully on a sample-by-sample basis even if the target amplicon doesn’t amplify.
Internal controls are produced by adding an additional primer pair that should always produce a control amplicon if template DNA is present. The control amplicon should be:
- significantly different in size than the target amplicon to allow easy visualisation of it and other amplicons
- larger than the target amplicon to reduce preferential amplification of the control amplicon, which could otherwise result in the target amplicon failing to amplify
For example, the gel results model below shows some expected results for a PCR assay for the PTC gene associated with bitterness tasting in humans. The assay should produce a control amplicon (366 bp in length) and either one or two smaller amplicons (260 and 151 bp). Two positive PCR controls (PC C/T and PC T/T) are used to ensure that the PCR assay is working. A positive control for C/C could also be included to make the PCR controls comprehensive.
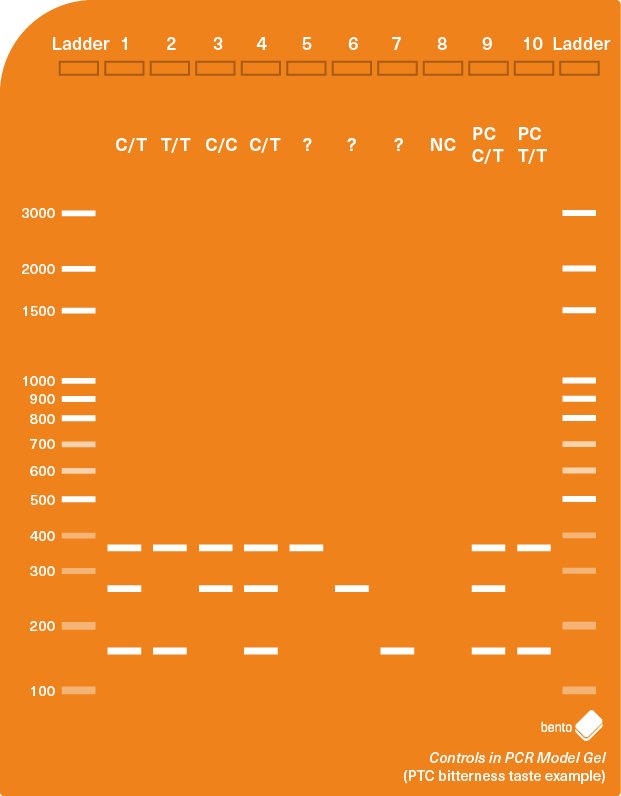
The inferences that you can make using an internal control in combination with negative and positive PCR controls are shown in the table below:
| Internal control | Target present | Positive PCR control | Negative PCR control | Inferences and next steps |
|---|---|---|---|---|
| Positive | Positive | Positive | Negative | Your PCR worked and the target DNA region is present. You have no systemic contamination. |
| Positive | Negative | Positive | Negative | Your PCR worked and your target DNA region is absent. You have no systemic contamination. |
| Negative | Positive | Positive | Negative | Your PCR worked and the target region is present. Your internal control is not working well. Try diluting your DNA extract. |
| Negative | Negative | Positive | Negative | Your sample PCR failed but the PCR worked in general. Troubleshoot the sample extraction. |
| Any result | Any result | Any failure of PCR controls | You should troubleshoot your PCR controls and repeat before inferring any results. | |
Conclusion
Controls are essential in most areas of scientific experimentation, and are especially important during DNA extraction and PCR to interpret results and to troubleshoot issues.
Controls allow you to establish a baseline for what should normally occur against which you can compare experimental results. They can be designed to test for many different possibilities by using known working examples of what you are experimenting with (for example DNA extracts or samples) to ensure you can control for unwanted variation and confounding factors.
By routinely using negative and positive PCR controls in your PCRs you will be able to assess your PCR results with increased confidence. You can also use multiple positive PCR controls to test whether multiplex PCR assays are working for multiple targets.
If you encounter PCR issues, you can then use DNA extraction controls to establish whether the problem is with the PCR, the DNA extraction, PCR inhibitors, or your samples themselves.
You can also use internal controls to make sure that PCRs are working on a sample by sample basis. This can be particularly important in assays where there is high variability of samples, or where the results of the assay are particularly important.
Please let us know what other resources, advice, and tips and tricks for using Bento Lab that you would like us to produce in the future!
Looking for advice on using Bento Lab?
Book a free consultation or ask a question.