Hi PCR enthusiasts!
Here are some recent highlights from Bento Lab.
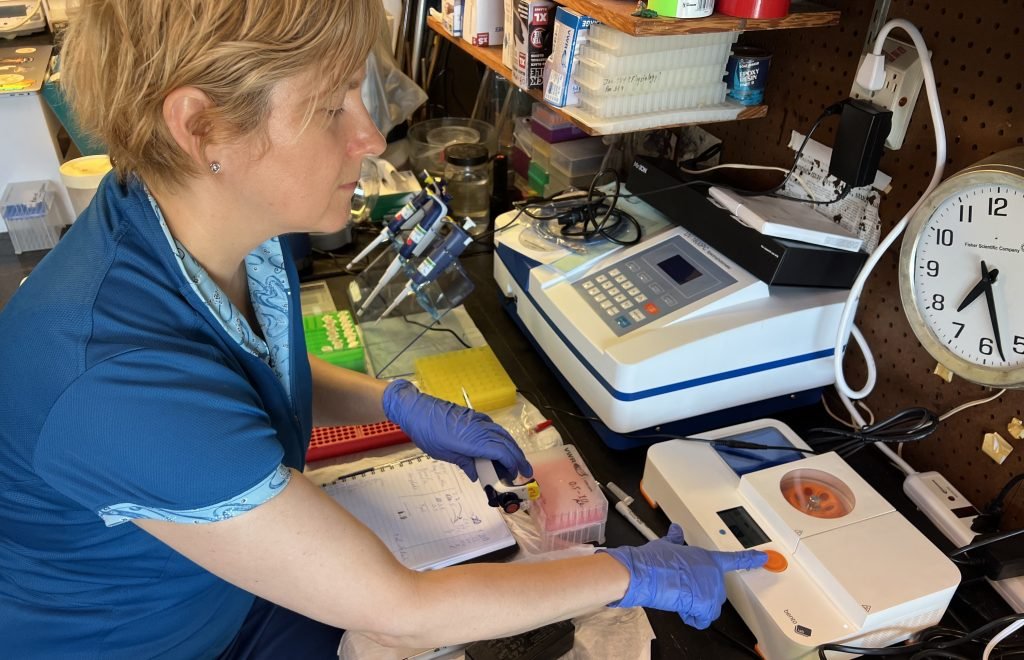
Bento Lab in the field
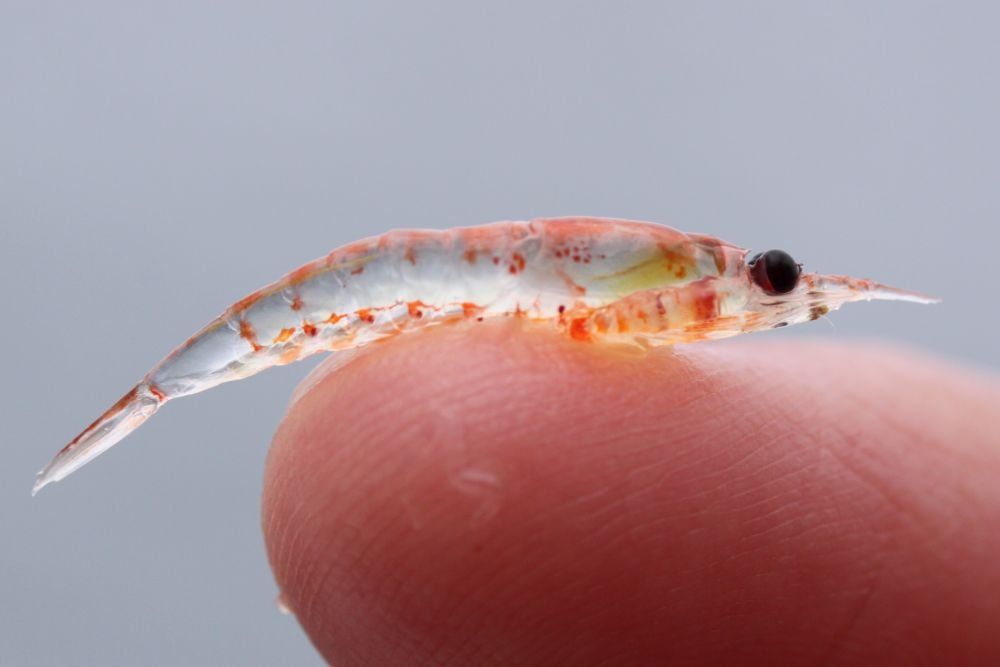
Unlocking the Mysteries of the Wild Ocean
We recently had the privilege of talking to Dr. Cheryl Ames, Professor of Applied Marine Biology, Tohoku University, Japan, about her exciting research.
Dr. Ames, an expert in jellyfish biodiversity and genomics, is making it possible to predict the presence of specific marine species in different regions of the ocean. Does today look like a cloudy day at the beach, with a high chance of deadly jellyfish?
Dr Ames wants to make such predictions a reality. She uses Bento Lab as a field PCR workstation and for quick and convenient troubleshooting in her lab.
Read the whole story about Dr. Ames’s awesome work here.
Interested in DNA methods and workflows? Subscribe for monthly insights.
Resources for Learners
We’ve been busy adding new articles to cover some of the obstacles faced by beginner (and even more experienced molecular biologists)
We would love to hear your feedback on these! Is anything good / bad, too long / too short, inspiring / boring, too simple / too complicated? Let us know how to support you better, and please feel free to send us your thoughts.
And if you have any particular requests for resources that would be useful to you or your colleagues, please get in touch.
Papers we love
This is a proof-of-concept article producing a detection workflow for a genus of plankton that causes Paralytic Shellfish Syndrome, one of the four recognised syndromes of shellfish poisoning in humans.
The authors, from the Centre for Environment Fisheries and Aquaculture Science (CEFAS), UK, came up with a number of novel and inventive approaches to doing this work in the field, including:
- Using a modified orbital power tool for cell lysis from sea water samples
- Using recombinase polymerase amplification instead of PCR to amplify DNA quickly without PCR inhibition
- Developing their own specific primers for genus Alexandrium
- Oxford Nanopore MinION metabarcoding
- Developing a portable power-efficient computer for nanopore read processing
- And much more!
Innovations at every step!
Bento Lab was used for incubations, PCR, and centrifugation in many places throughout their workflows.
You can read the entire paper here.
Methods and Techniques: PCR-RFLP
We were inspired to take a look at PCR-RFLP by a recently published five-year study of mammal biodiversity in a reserve in Moldova (Nistreanu et al. 2023, from a team associated with the Institutul de Zoologie, Universitatea de Stat din Moldova).
As a small part of this study, the researchers used Bento Lab to distinguish between wild boars from feral pigs using a PCR-RFLP assay (PCR followed by restriction enzyme digests and gel electrophoresis). Read the paper here (it’s mostly written in Romanian) and if you’re interested in the protocol you can check out the article where it was first published.
RFLP (restriction fragment length polymorphism) is a “DNA fingerprinting” technique whereby a DNA region is amplified and cut using enzymes at a specific nucleotide motif. The pattern of fragment sizes can be used to characterise and/or identify the sample when analysed by gel or capillary electrophoresis.
Wikipedia claims that RFLP is a “largely obsolete” method, but we were able to find a wide range of PCR-RFLP applications used as recently as 2023, such as in bird sexing; parasite detection; identifying different animals (such as ghost crabs in this article); rapid screening of fungal cultures; differentiating coffees of different origins, and authenticating grape varieties used in wine.
We think that PCR-RFLP could be a powerful tool for people wanting to do similar work to the above with Bento Lab, as a step up from conventional PCR assays without requiring the additional costs of DNA sequencing.
What do you think?
Looking for advice on using Bento Lab?
Book a free consultation or ask a question.